The Data Repository of Antiviral peptides and proteins (DRAVP) was created for the purpose of helping users browse and retrieve AVPs in a more convenient way. The database is manually curated and contains information of AVPs from journals, patents and clinical trials platforms.
Help
Antiviral peptide
General dataset is the most important part of DRAVP because each entry has detailed annotations and serious descriptions.
| - | Fields | Description |
|---|---|---|
| General information | DRAVP ID | The field provides the unique accessing number linking to the corresponding DRAVP entry |
| Peptide Name | Name of each peptide or protein in DRAVP. | |
| Sequence | The peptide sequence which is represented by single letter codes. | |
| Sequence Length | Number of resiudes in the peptide sequence | |
| Uniprot ID | Provide the accessing link(s) directing to external Uniprot entry(or entries). | |
| Source | The organism where the peptides or proteins were extracted or isolated. | |
| Activity information | Target Organism | Specific virus types |
| Assay | Methods for determination of antiviral activity in literature | |
| Activity | Antiviral activity information against viruses | |
| Hemolytic Activity | Hemolytic activity information against red blood cells (RBCs) | |
| Cytotoxicity | cytotoxicity information against host cells except RBCs | |
| Binding Target | The action site of peptides against viruses | |
| Mechanism | The mechanism of peptides exhibit antiviral activity | |
| Structure information | PDB ID | Provide accessing link(s) directing to the correspong PDB entry. |
| Predicted Structure ID | Structure predicted by Alphafold,click can download the PDB files | |
| Structure | Show the structure with Mol*viewer | |
| Linear/Cyclic | Linear or cyclic of peptides | |
| N/C-terminal Modification | The modifications of N/C-terminal according to the references | |
| Other Modification | All bonds and special amino acids (out of 20 common amino acids) | |
| Stereochemistry | The L/D amino acids consist peptides | |
| Physicochemical Information | Formula, mass, pI, Net charge and other information. | |
| Literature Information | The information of peptides come from all kinds of literature and the section provides the way to find the full text. | |
Antiviral protein
Proteins with antiviral activity have been collected in the dataset. The annotation information of protein data is similar to that of general data, but there is no physicochemical information. The standard of collection of proteins are as follow:
1. Sequence length ≥ 100.
2. Direct or indirect antiviral effects are specifically reported in the literature.
Patent Data
Patent dataset is based on a large amount of patent sequences, which account for a large proportion in DRAVP. Such patent AVPs information can show existing patented AVP sequences and help researchers avoid infringement risk. The page of patent AVPs present patent accession number and family information. Users can browse detailed patent information in Lens.org.
Clinical Data
Clinical dataset is an important part of our database, although the clinical data are fewer comparing to other datasets. We have collected the clinical trials information in DRAVP, which contains NCT Number, Condition/Disease, phase and Study Title.
Specific Data
Specific dataset contains stapled AVPs.Stapled AVPs are a special subset of antiviral peptides that contain hydrocarbon staples or other chemical bridges to stabilize α-helical conformations. These modifications often enhance proteolytic stability, cell permeability and antiviral potency. Each entry includes stapling positions, cross-linker chemistry and structural annotations.
Simple search
The simple search page allows users to search individual fields found within AVP.
1. Find a list of all indexed fields in the drop-down menu and choose one of your interest.
2. Enter the appropriate contents in the text area below.
3. Click "Submit" (or click "Reset" to clear your input).

- Sequence >>> Single letter code (no space, mature peptide only) .e.g.INLKAIAALAKKLL
- Peptide Name >>> Name of peptides in DRAVP (full name or short name works) .e.g.Mastoparan M
- Sequence length >>> Enter the peptide sequence length number. e.g.21
- Target Organism >>> The virus type which AVP inhibits e.g.HIV.
- Linear/Cyclic >>> Structural properties of peptides, linear or cyclic. e.g.linear
- UniProt ID >>> Accessing number and linking to UniProtKB/Swiss-Prot entries. e.g.P04205
- PDB ID >>> Accessing numble of Protein Data Bank. e.g.2KHT
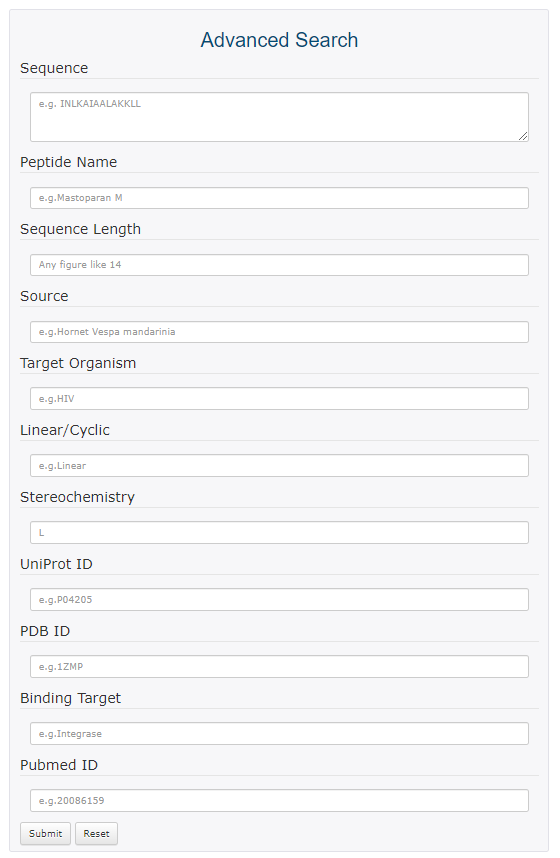
Advanced search
Enter the appropriate contents in the text area. The relationship between each item is "and".
Blast
The BLAST (Basic Local Alignment Search Tool) program uses a strategy based on matching sequence fragments by employing a powerful statistical model to find the best local alignments (For more information see http://www.ebi.ac.uk/Tools/sss/ncbiblast/).
Usage Introduction
1. Sequence Input: The query sequence can be entered directly into text area. The sequence must be FASTA format.
2. Parameter choose: choose the scoring matrix to be applied to the search, the default value is BLOSUM62.
